Rescale で PETSc アルゴリズムのスケーラビリティをテストする

科学計算用ポータブル拡張ツールキット (PETSc) は、偏微分方程式によってモデル化された科学アプリケーションのスケーラブルな (並列) ソリューションのためにアルゴンヌ国立研究所によって開発されたデータ構造とルーチンのスイートです。 PETSc は、偏微分方程式および疎行列計算用に世界で最も広く使用されている並列数値ソフトウェア ライブラリの XNUMX つです。
従来、科学者やアルゴリズム開発者が PETSc で新しい並列アルゴリズムを完成させると、それをマルチコア コンピューター クラスターで実行して、そのスケーラビリティと高速化をテストする必要がありました。 クラスターは通常、HPC リソースを維持するために多大な管理作業を伴う大学、政府、または企業の共有コンピューティング リソースです。 さらに、実行を作成するには、科学者または開発者は環境を準備する必要がありますが、これは困難で時間がかかる可能性があり、実行中に予期しないことが発生するとエラーが発生し、出力データが生成されない可能性があります。
Rescale を使用すると、PETSc アルゴリズムのスケーラビリティのテストがはるかに簡単になります。 科学者または開発者は、ハードウェアのタイプとコアの数を指定し、インターネット接続と Web ブラウザを使用してジョブを実行できます。
テストするアルゴリズム
私がテストしようとしているアルゴリズムは、PETSc パッケージの公式チュートリアルからのものです。 このコードは、KSP と並行して線形システムを解きます。 KSP を選択したのは、KSP が PETSc パッケージで最もよく使用される操作の XNUMX つであるためです。 さらなる分析のために、すべてのプロセスの各 PETSc 関数呼び出しの操作のタイムスタンプをログ ファイルに出力するように若干の変更を加えました。
ここにある ソースコード 私の修正後。 私も作成しました メイクファイル それをコンパイルするためです。
KSP が行うことは、n 個の要素を持つベクトル X に対して線形方程式 AX=B を解くことです。ここで、A は mxn サイズの行列、B は m 個の要素を持つベクトルです。
Rescale でアルゴリズムを実行する
Rescale にサインアップすると、PETSc アルゴリズムをコンパイルして実行できる新しいジョブを作成できます。
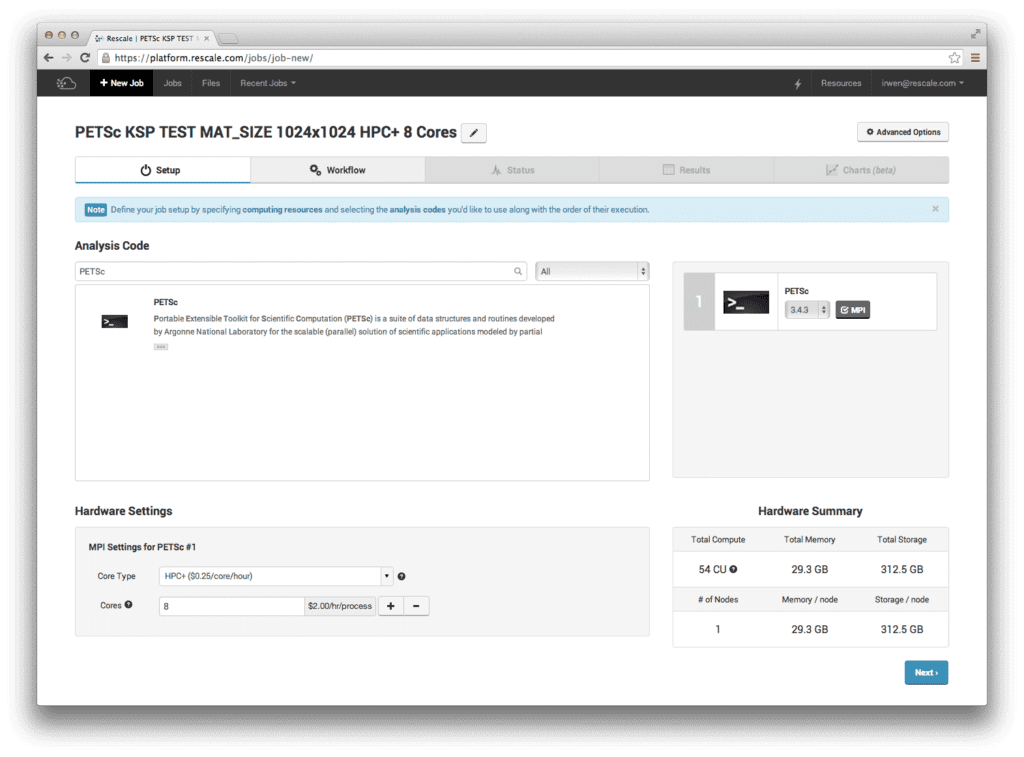
[セットアップ] ページで、[分析コード] セクションから PETSc を選択します。 [ハードウェア設定] で、[コア タイプ] として 8 コアの HPC+ を選択します。 下の画像は、画面がどのように見えるかを示しています。
[ワークフロー] ページで、ソース コードとメイクファイルをアップロードします。 代わりに、コンパイルされた実行可能バイナリ ファイルをアップロードすることも選択できます。 分析コマンドに実行したいコマンドを入力します。
作る; mpirun -n 8 ./ksp_test -n 1024 -m 1024 -logfile ksp_test_1024_1024_8.log
[ワークフロー] ページは次のようになります。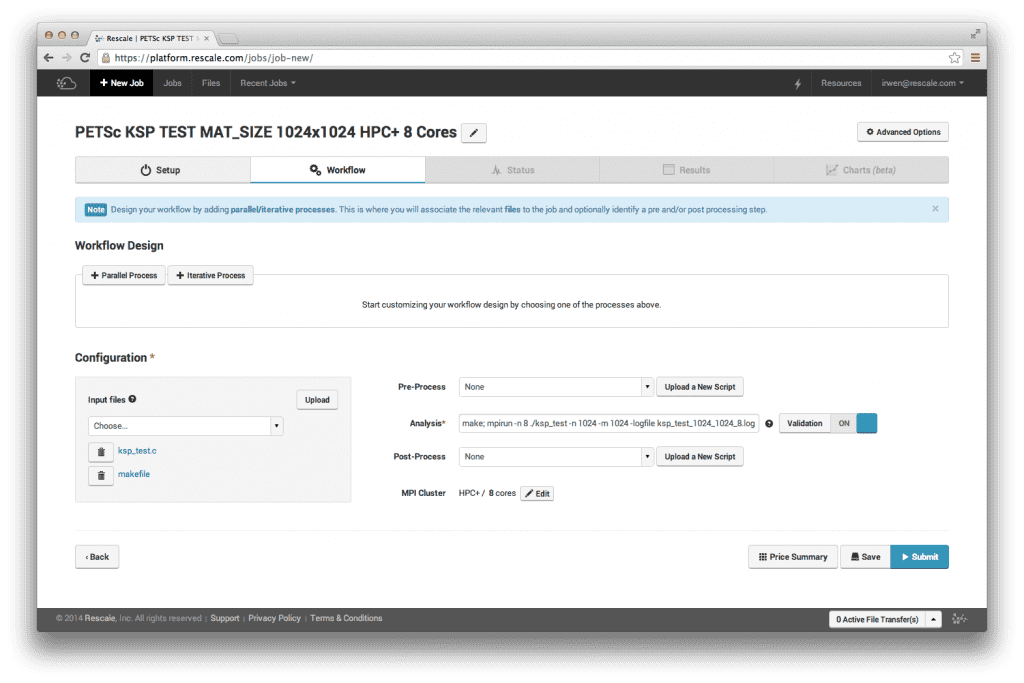
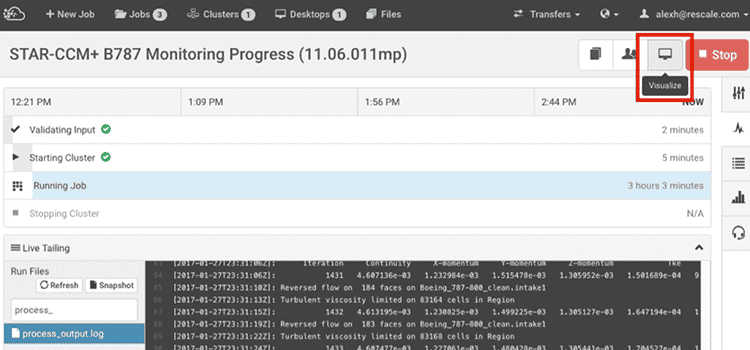
右下隅にある「送信」をクリックして、ジョブを実行して再スケールします。 ジョブを送信した後、[ステータス] ページでジョブをリアルタイムで監視できるようになります。
ジョブが完了すると、結果ページから出力ファイルとログ ファイルを表示およびダウンロードできます。
KSP テスト結果
スケーラビリティ テストでは、Rescale の HPC+ コア タイプで 1、2、4、8、16、および 32 コアを選択しました。 行列 A のサイズは 1024 x 1024 でした。平均プロセス実行時間、反復回数、および反復あたりの時間の結果は次のとおりです。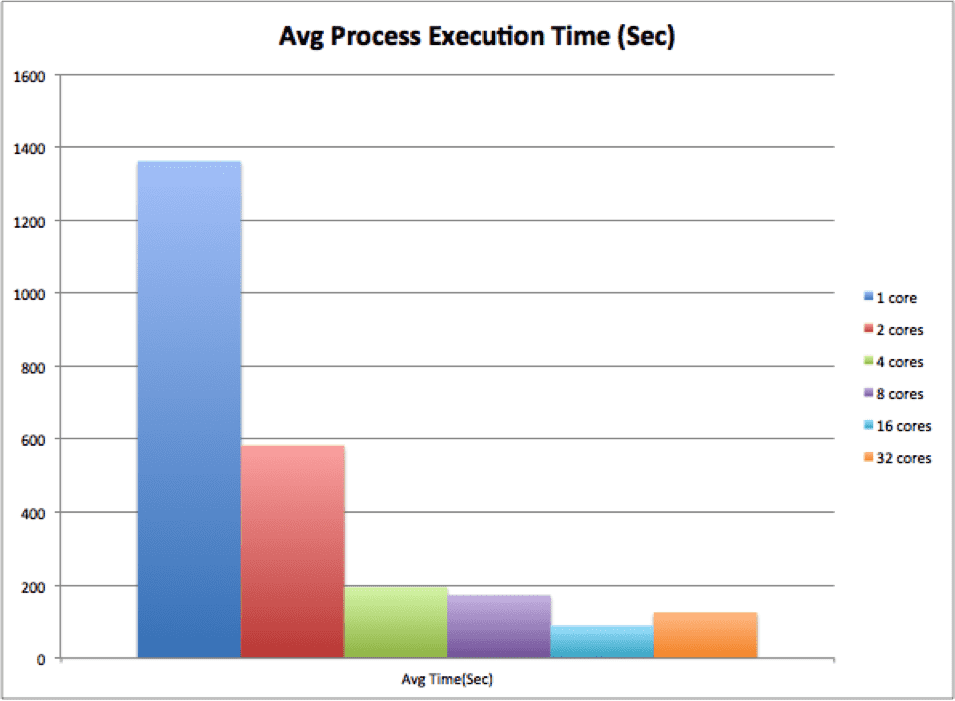
平均プロセス実行時間から、コア数が増加するにつれて時間が減少し (最大 16 コアまで)、その後 32 コアになると予想外に増加したことがわかります。 これは、反復回数を考慮する必要があるために発生しました。
アルゴリズムが開始されるたびに、行列と右手ベクトルがランダムに生成されます。 これは、「誤差の基準」に収束するために必要な反復が毎回異なることを意味します。 次のグラフでは、テストの各実行の反復数が表示されます。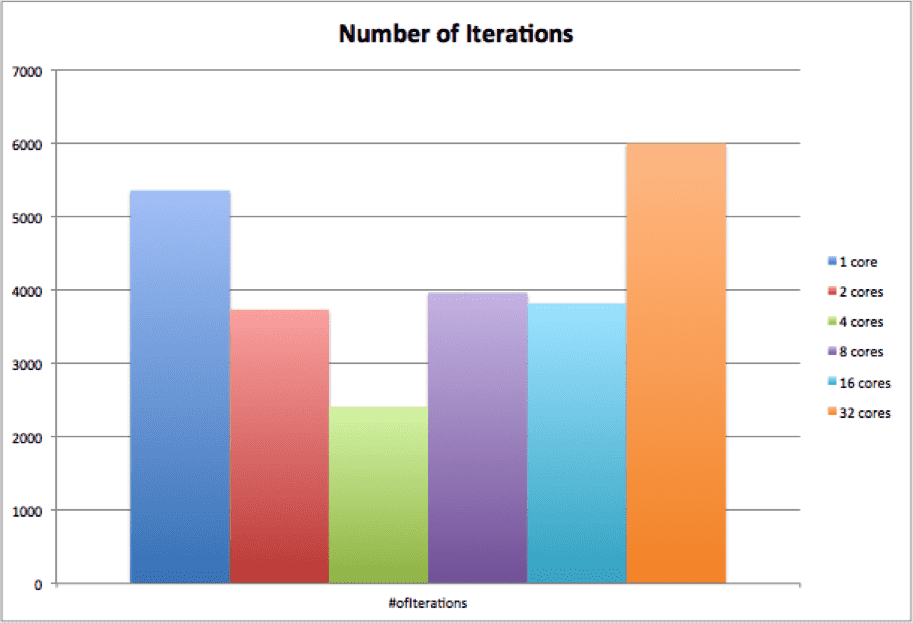
最後のグラフは、XNUMX 秒あたりの反復数を示します。これは、反復数 / 平均プロセス実行時間で計算されます。 グラフから、並列 KSP はコア数の増加に合わせて適切に拡張できることがわかります。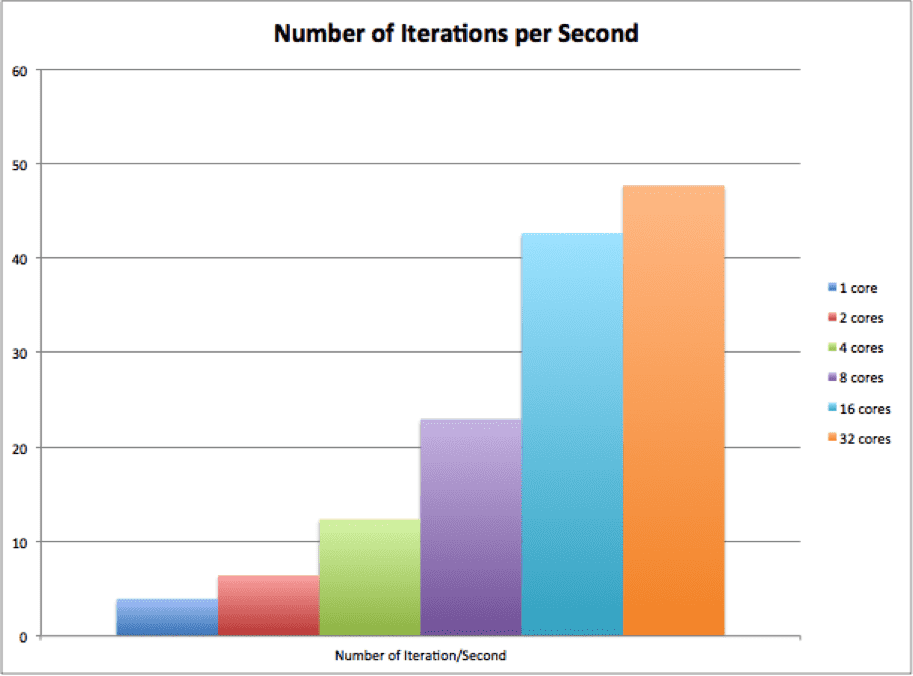
Rescale アカウントをお持ちの場合は、 こちら 先ほど述べた KSP テスト ジョブのクローンを作成し、異なるハードウェア設定、コア数、パラメーターを使用してシミュレーションを実行してみます。 をクリックすることもできます こちら 「PETSc helloworld」サンプル ジョブのクローンを作成します。 アカウントをお持ちでない場合は、クリックしてください こちら サインアップします。